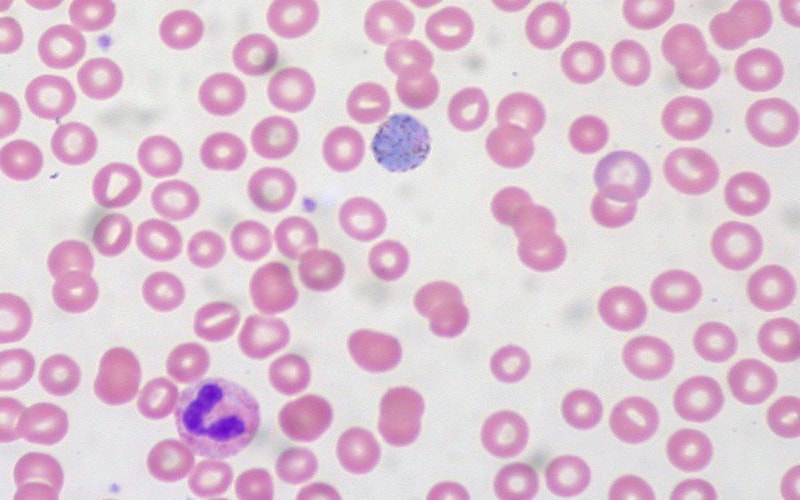
Sendai: Researchers have developed a robust network link map on malaria transactions between human host cells.
Malaria parasites transform healthy red blood cells into rigid versions of themselves that clump together, hindering the transportation of oxygen.
According to the Malaria Report 2018 of the World Health Organization, the infectious disease affects more than 200 million people worldwide and causes up to half a million deaths each year.
The research study was published in iScience, a Cell Press journal.
Plasmodium falciparum was the most serious malaria-causing parasite of the researchers. This virus infects a red blood cell of the host, and induces the production in the cytoplasm of the host cell of several proteins– the bulk of its structure and the fluid in which it is stored, thereby altering the physical shape of the tissue.
It not only lets the cells adhere, but it also allows the virus to move to the cell’s surface to invade them, from the body’s immune response. Together, proteins act to replicate the parasite, causing the malaria virus to spread.
Kentaro Kato, a professor at Tohoku University Graduate School of Agricultural Science said: “Our study sheds light on the highly complicated interplay between parasite and host proteins in the host cytoplasm. The work provides a reliable dataset of the interactions connecting dozens of proteins the parasite exports to continue infecting the host cells.”
Previous to this, because the Parasite was expected to export about 400 proteins, it was difficult to understand how the parasite works with the activated proteins. Protein without the genetic sequence could also be released into the cytoplasm of a cell.
In this study, the researchers opted to focus on one of these proteins without the parasitic mark–skeleton-binding protein 1 (SBP1), which is known to be highly important for malaria to propagate.
By studying a protein known to be related to malaria virulence, but that isn’t specifically triggered by the parasitic proteins, the researchers could narrow in on specific protein interactions to understand how the infection traffics within and beyond the host cells.
During the entire proliferation process, highly sensitive mass spectrometry was used to image proteins that interact with SBP1 and thus to identify several proteins specifically linked to host cell transformation.
“In this study, we developed an alternative approach to identify exported proteins involved in the trafficking complex and in the parasite protein exports. The SBP1 interactions established in our study represent a powerful and invaluable platform to identify exported proteins related to severe malaria caused by Plasmodium falciparum,” Kato added.
Research provided an extensive map of SBP1 interactions, which illuminated the complex relationships between host and parasite protein and the interaction between them.